 
Early results on simple model systems showed that ML could provide highly accurate values for T s with only modest amounts of training 20, but that the corresponding functional derivatives are too noisy to yield sufficiently accurate results to (i). 
The solution of the KS equations performs both tasks exactly. 1: (i) its functional derivative is used in the Euler equation which is solved in the self-consistent cycle and (ii) when self-consistency is reached, the ground-state energy of the system is calculated by E, an orbital-free (OF) mapping. The non-interacting kinetic energy functional T s of the electron density n is used in two distinct ways in a KS calculation 1, as illustrated in Fig. If such attempts could be made practical, the possible speed-up in repeated DFT calculations of similar species, such as occur in ab initio MD simulations, is enormous.Ī key difficulty has been the need to extract the functional derivative of the non-interacting kinetic energy.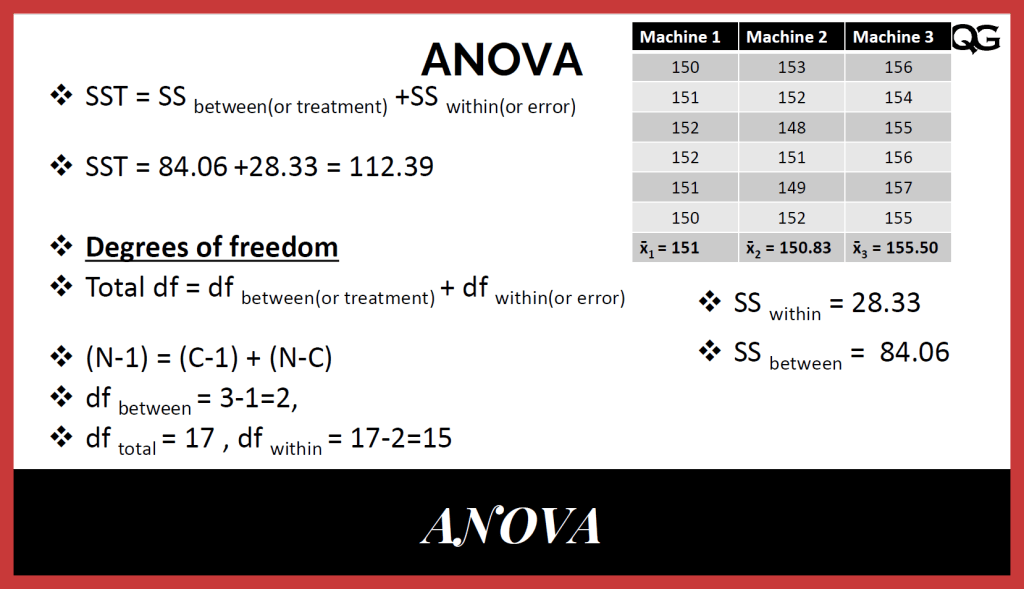 
Fewer still have focussed on finding the functionals of DFT as a method of performing KS electronic structure calculations without solving the KS equations 20, 21, 22, 23, 24. A few applications involve finding potential energy surfaces within molecular dynamics (MD) simulations 16, 17, 18, 19. The majority of these applications involve predicting properties of molecules or materials from large databases of KS-DFT calculations 12, 13, 14, 15. There has also been a recent spike of interest in applying machine learning (ML) methods in the physical sciences 7, 8, 9, 10, 11. Such calculations are playing a key role in the materials genome initiative 5, at least for weakly correlated materials 6. Useful accuracy is achieved with standard exchange–correlation (XC) approximations, such as generalized gradient approximations 3 and hybrids 4. Kohn–Sham density functional theory 1 (KS-DFT) is now enormously popular as an electronic structure method in a wide variety of fields 2. Learning density models now allows the construction of accurate density functionals for realistic molecular systems. We perform the first molecular dynamics simulation with a machine-learned density functional on malonaldehyde and are able to capture the intramolecular proton transfer process. The present work overcomes this difficulty by directly learning the density-potential and energy-density maps for test systems and various molecules. This should yield substantial savings in computer time, allowing larger systems and/or longer time-scales to be tackled, but attempts to machine-learn this functional have been limited by the need to find its derivative. Machine learning holds the promise of learning the energy functional via examples, bypassing the need to solve the Kohn–Sham equations. Subsample a subset of \(s\) variables, \(\mathcal\), the \(p\)-value is adjusted using a multiple comparison method such as the Bonferroni method or the Benjamini-Hochberg procedure.Last year, at least 30,000 scientific papers used the Kohn–Sham scheme of density functional theory to solve electronic structure problems in a wide variety of scientific fields. We let \(m\) be the number of subsamplings, \(s\) be the sampling size, \(q\) be both the number of subsamplings to keep, and the number of semifinalist variables. Let \(d\) be the (total) dimension and \(d_0\) be the number of true variables. It now captured X2,X3,X4,X5, and also a noise variable X6 that is not as significant as the former 4 covariates X2-X4. Residual standard error: 1.371 on 94 degrees of freedom :max_bytes(150000):strip_icc()/degrees-of-freedom-c87f4f95d13043da80a0735bd773d948.jpg)
The stepwise selection removes X1, which has the weakest signal (hence is not expected to be recovered.) We next check the results with a larger m = 1000. The final model perfectly recovers the five true variables. Residual standard error: 1.372 on 95 degrees of freedom Lm(formula = yy ~ X5 + X3 + X2 + X4, data = dataXY) Residual standard error: 1.375 on 94 degrees of freedom 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
